Documentation of eeglib.helpers module¶
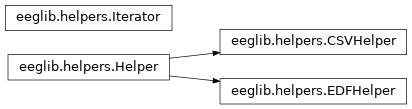
This module contains helper classes that are useful to iterating over a EEG data stream. Currently there is support only for CSV files.
- class eeglib.helpers.CSVHelper(path, *args, **kargs)¶
Bases:
HelperThis class is for applying diferents operations using the EEG class over a csv file.
Methods
getNames([indexes])Returns the names of the specified indexes of channels.
moveEEGWindow(position)Moves the window to start at position.
prepareEEG(windowSize)Prepares and creates the EEG object that the iteration will use with the same parameters that an EEG objects is initialized.
prepareIterator([step, startPoint, endPoint])Prepares the iterator of the helper.
selectSignals(selectedSignals)- Parameters
- __init__(path, *args, **kargs)¶
The rest of parameters can be seen at
Helper.__init__()- Parameters
- path: str
The path to the csv file
- getNames(indexes=None)¶
Returns the names of the specified indexes of channels.
- Parameters
- indexes: Iterable of int, optional.
The indexes of the channels desired. If None it will return all the channels’ names. Default: None.
- moveEEGWindow(position)¶
Moves the window to start at position. Also it returns the inner eeg object.
- Parameters
- position: int, str or datetime.timedelta
int: position of the sample
str with format hh:mm:ss.ss: temporal position
timedelta: temporal position
- Returns
- EEG
- prepareEEG(windowSize)¶
Prepares and creates the EEG object that the iteration will use with the same parameters that an EEG objects is initialized. Also it returns the inner eeg object.
- Parameters
- windowSize: int
The maximun samples the window will store.
- Returns
- EEG
- prepareIterator(step=None, startPoint=0, endPoint=None)¶
Prepares the iterator of the helper.
- Parameters
- step: int,optional
Number of samples to be skipped in each iteration.
- startPoint: int, optional
The index of first sample from where the iteration will start. By default 0.
- endPoint: int, optional
The index of the last sample + 1 until where the iteration will go. By default the size of the data.
- selectSignals(selectedSignals)¶
- Parameters
- selectedSignalsiterable of str and/or int
The indexes of the desired selected signals. It must be an iterable type containing either str or int.
- Returns
- None.
- class eeglib.helpers.EDFHelper(path, *args, sampleRate=None, **kargs)¶
Bases:
HelperThis class is for applying diferents operations using the EEG class over an edf file.
Methods
getNames([indexes])Returns the names of the specified indexes of channels.
moveEEGWindow(position)Moves the window to start at position.
prepareEEG(windowSize)Prepares and creates the EEG object that the iteration will use with the same parameters that an EEG objects is initialized.
prepareIterator([step, startPoint, endPoint])Prepares the iterator of the helper.
selectSignals(selectedSignals)- Parameters
- __init__(path, *args, sampleRate=None, **kargs)¶
The rest of parameters can be seen at
Helper.__init__()- Parameters
- path: str
The path to the edf file
- getNames(indexes=None)¶
Returns the names of the specified indexes of channels.
- Parameters
- indexes: Iterable of int, optional.
The indexes of the channels desired. If None it will return all the channels’ names. Default: None.
- moveEEGWindow(position)¶
Moves the window to start at position. Also it returns the inner eeg object.
- Parameters
- position: int, str or datetime.timedelta
int: position of the sample
str with format hh:mm:ss.ss: temporal position
timedelta: temporal position
- Returns
- EEG
- prepareEEG(windowSize)¶
Prepares and creates the EEG object that the iteration will use with the same parameters that an EEG objects is initialized. Also it returns the inner eeg object.
- Parameters
- windowSize: int
The maximun samples the window will store.
- Returns
- EEG
- prepareIterator(step=None, startPoint=0, endPoint=None)¶
Prepares the iterator of the helper.
- Parameters
- step: int,optional
Number of samples to be skipped in each iteration.
- startPoint: int, optional
The index of first sample from where the iteration will start. By default 0.
- endPoint: int, optional
The index of the last sample + 1 until where the iteration will go. By default the size of the data.
- selectSignals(selectedSignals)¶
- Parameters
- selectedSignalsiterable of str and/or int
The indexes of the desired selected signals. It must be an iterable type containing either str or int.
- Returns
- None.
- class eeglib.helpers.Helper(data, sampleRate=None, windowSize=None, names=None, highpass=None, lowpass=None, normalize=False, ICA=False, selectedSignals=None)¶
Bases:
objectThis is an abstract class that defines the way every helper works.
Methods
getNames([indexes])Returns the names of the specified indexes of channels.
moveEEGWindow(position)Moves the window to start at position.
prepareEEG(windowSize)Prepares and creates the EEG object that the iteration will use with the same parameters that an EEG objects is initialized.
prepareIterator([step, startPoint, endPoint])Prepares the iterator of the helper.
selectSignals(selectedSignals)- Parameters
- __init__(data, sampleRate=None, windowSize=None, names=None, highpass=None, lowpass=None, normalize=False, ICA=False, selectedSignals=None)¶
- Parameters
- data: 2D matrix
The signals data in the shape (nChannels, nSamples).
- sampleRate: numeric, optional
The frequency at which the data was recorded. By default its value is the lenght of the data.
- windowSize: int, optional
The size of the window in which the calculations will be done. By default its value is the lenght of one second of the data.
- names: list of strings
A list containing the names of each channel in the same positions than data channels.
- highpass: numeric, optional
The signal will be filtered above this value.
- lowpass: numeric, optional
The signal will be filtered bellow this value.
- normalize: boolean, optional
If True, the data will be normalizing using z-scores. Default = False.
- ICA: boolean, optional
If True, Independent Component Analysis will be applied to the data . It is applied always after normalization if normalize = True. Default: False.
- selectedSignals: list of strings or ints
If the data file has names asociated to each columns, those columns can be selected through the name or the index of the column. If the data file hasn’t names in the columns, they can be selected just by the index.
- getNames(indexes=None)¶
Returns the names of the specified indexes of channels.
- Parameters
- indexes: Iterable of int, optional.
The indexes of the channels desired. If None it will return all the channels’ names. Default: None.
- moveEEGWindow(position)¶
Moves the window to start at position. Also it returns the inner eeg object.
- Parameters
- position: int, str or datetime.timedelta
int: position of the sample
str with format hh:mm:ss.ss: temporal position
timedelta: temporal position
- Returns
- EEG
- prepareEEG(windowSize)¶
Prepares and creates the EEG object that the iteration will use with the same parameters that an EEG objects is initialized. Also it returns the inner eeg object.
- Parameters
- windowSize: int
The maximun samples the window will store.
- Returns
- EEG
- prepareIterator(step=None, startPoint=0, endPoint=None)¶
Prepares the iterator of the helper.
- Parameters
- step: int,optional
Number of samples to be skipped in each iteration.
- startPoint: int, optional
The index of first sample from where the iteration will start. By default 0.
- endPoint: int, optional
The index of the last sample + 1 until where the iteration will go. By default the size of the data.
- selectSignals(selectedSignals)¶
- Parameters
- selectedSignalsiterable of str and/or int
The indexes of the desired selected signals. It must be an iterable type containing either str or int.
- Returns
- None.